3 Intestinal Development: Structural Mechanisms
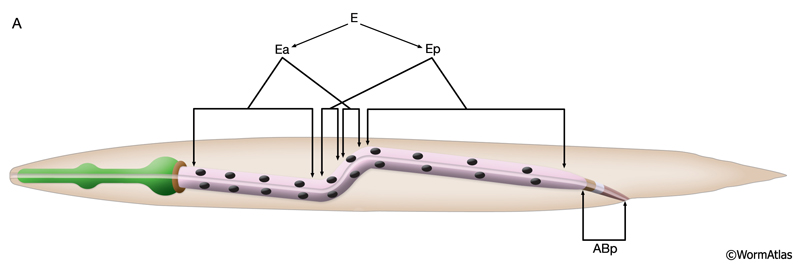
The E blastomere is born on the surface of the embryo at about 35 min after fertilization (IntFIG 3). From this point on, the specific stages of intestinal development are indicated by the number of E descendants present such as E2, E4, E8, E16 and E20, although occasionally due to an extra cell division during development the mature intestine is seen to be made of 21 cells instead of the usual 20 (Sulston and Horvitz, 1977). The daughters of E, E.a and E.p, migrate into the interior of the embryo initiating gastrulation when the embryo is at the 26-cell stage (Bucher and Seydoux, 1994, Nance et al., 2005). At the E16 stage, the intestinal primordium has a ventral tier of six cells and a dorsal tier of ten cells (IntFIG 4). About 30 min into the E16 stage, cytoplasmic polarization of intestinal cells occurs such that the nuclei of cells move towards and cytoplasmic components move away from the midline (Leung et al., 1999). Shortly afterwards, cell separation starts at the midline as small gaps, and these small gaps eventually become the lumen of the intestine. At the same time, electron-dense vesicles begin to appear in the cytoplasm and localize near the basal pole. These vesicles may correspond to the intestine-specific gut granules (IntFIG 1). At E16-E20, two ventral cell pairs intercalate between the dorsal cells, resulting in a single layer of intestinal cells with bilateral symmetry. As the second cell intercalation occurs, neighboring int II, int III and int IV rings initiate a coordinated 90° clockwise rotation around the axis of the midline. Between 430 min following first cell division and hatching, intestinal rings VII-IX make coordinated 90° counter-clockwise rotation which leads to the twisted appearance of the intestine in the newly hatched larva (IntFIG 3) (Sulston and Horvitz, 1977; Mendenhall et al., 2015). As a result, int 5L connects to int 4V and int 6L connects to int 7L in the adjoining rings (Mendenhall et al., 2015; A. Mendenhall, pers. comm.; Z. F. Altun and D. H. Hall, unpub. observations). The cells in int V and/or int VI rings get variably pushed to the right or left side by the developing uterus in subsequent stages and, hence, their nuclei are stochastically located to the right or left of the midline. The forces or developmental processes that influence the positions of these cells in postembryonic life are still unknown. By the adult stage, the intestine is composed of 20 cells with a total of 30-34 nuclei and 32C per nucleus (IntFIG 4X).These cell movements result in a superhelical twist of the intestine, displacing the anterior half to the left side of the larval body and the posterior half to the right side. This twist of the intestine, in turn, is suggested to lead to the asymmetrical growth of the gonad later in life (Hermann et al., 2000). The left-right rotational asymmetry of this twist is determined by the LIN-12/Notch pathway and involves LAG-2, APX-1 and LAG-1 proteins. Also a pathway involving POP-1 and LIT-1 limits this twist to the anterior half of the intestine. Subsequently, the intestinal primordium elongates (Hermann et al., 2000). By the time of hatching, the anterior intestinal rings may make an additional 90° rotation (IntFIG 2 and IntFIG 3) (Sulston and Horvitz, 1977). This second turn of the anterior intestinal cells seems to be variable, however, because cells in int II-IV rings can often be seen as dorsoventral to each other in adult animals. Similarly, orientation of cells in the adult int VI-IX rings is variable. Rings VI-VIII tend to adopt L/R positions, whereas ring IX cells are usually positioned dorsoventrally (Z.F. Altun and D.H. Hall, unpubl.).
IntFIG 4B-J: Development of the intestine from E8 to L2 stages. Anterior is to the left. (B-D) DIC images. (E-G) Epifluorescent images of transgenic animals expressing the reporter gene pW02H5.8-NLS::GFP. (H-J) Merged DIC and epifluorescent images. (Pink dots) Basal lamina, detected on the intestinal surface beginning in early morphogenesis. (Strain source: M. Molin, A. Blomberg and M. Pilon.) Magnification, 600x.
IntFIG 4K-W: Development of the intestine from E8 to L2 stages. Anterior is to the left. (K-M, T and V) DIC images. (N-P, U and W) Epifluorescent images of transgenic animals expressing the reporter gene pW02H5.8-NLS::GFP. (Q-S) Merged DIC and epifluorescent images. (Pink dots) Basal lamina, detected on the intestinal surface beginning in early morphogenesis. (Strain source: M. Molin, A. Blomberg and M. Pilon.) Magnification, 600x.
IntFIG 4X: Endoreduplication of intestinal nuclei. (Based on Sulston and Horvitz 1977; Hedgecock and White 1985; Leung et al., 1999.)
During epithelial polarization, which follows cell intercalation, punctate foci of adherens junction proteins organize into rectilinear junctions surrounding the lumen of the intestine. Through this process, each cell acquires distinctive apical and basal surfaces. During subsequent embryogenesis, the apical membranes of cells between the adherens junctions increase greatly in area as microvilli develop, and correspondingly, the apical surface of the intestine expands. In addition, later, the cytoplasmic polarization disappears so that intestinal nuclei are found in the center of the cells and other organelles are more evenly dispersed within the cytoplasm (Leung et al, 1999).
Intestinal cells become binucleate and polyploid during post-embryonic development (Hedgecock and White, 1985). At the beginning of the lethargus of the first molt, most of the intestinal nuclei, except the anteriormost 6, divide without accompanied cell divisions giving rise to 20 intestinal cells with a total of 30-34 nuclei. Despite a large increase in tissue volume, the intestine continues to grow without further cell or nuclei divisions. Intestinal nuclei continue to increase in size and go through repeated endoreduplications (chromosome duplication without karyokinesis), increasing the ploidy of each nucleus to 32C by the final molt. These endoreduplications are generally synchronized to each period of lethargus, resulting in a twofold increase in chromosomal number at the end of each molt. By the adult stage, the intestine is composed of 20 cells with a total of 30-34 nuclei and 32C per nucleus.
|
4 Intestine Structure and Function
The intestine is composed of large, cuboidal cells, with distinct apical, lateral and basal regions (IntFIG 5) (see also Gap Junctions). Each intestinal cell forms part of the intestinal lumen at its apical pole and secretes the constituents of the basal lamina from its basal pole. The intestinal basal lamina contains laminin α and β nidogen/entactin, which are made by the intestine, and type IV collagen, which is made by the muscle and somatic gonad (Graham et al., 1997; Kang and Kramer, 2000; Huang et al., 2003; Kao et al., 2006). Each intestinal cell is sealed laterally to its neighbors by large adherens junctions close to the apical side (Labousse, 2006). It also connects to the neighboring intestinal cells via gap junctions on the lateral sides (IntFIG 5). The lateral membranes also display a region of tightly folded plasma membranes that may represent another specialized intercellular junction of novel form.
IntFIG 5A & B: Ultrastructure of the intestinal cells. Pseudocolored, transverse TEM images. (DC) Dorsal cord; (VNC) ventral cord.
A. Section thorough the pharyngeal intestinal valve cells (brown). (Inset) Lengthwise view. Bar, 1 μm. (Image source: [Hall] N510-R338.
B. Section thorough the first intestinal ring, composed of four cells. (Image source: [Hall] N510-R333.)
IntFIG 5C-F: Ultrastructure of the intestinal cells. Pseudocolored, transverse TEM images. (DC) Dorsal cord; (VNC) ventral cord. C. Section through the anterior body. Bar. 1 μm. (Image source: [Hall] N510-G127.) D. Section through the last intestinal ring. (PAG) Preanal ganglion. (Image source: [Hall] B140-T579.) E. Intestinal lumen of the third intestinal ring, magnified. Same image as in C. A complex junction (arrow) next to an adherens junction (arrowhead) seals the two intestinal cells to each other (inset). A basal lamina covers the surface of the intestine facing the pseudocoelom. F. Diagram of a cross section through an intestinal cell.
Many microvilli extend into the lumen from the apical face, forming a brush border (IntFIG 5). The microvilli are anchored into a strong cytoskeletal network of intermediate filaments at their base, called the terminal web. The core of each microvillus has a bundle of actin filaments that connects to this web. Over the microvilli, there is an extracellular electron-lucent coating of highly modified glycoproteins (a glycocalyx), which may function to localize digestive enzymes, protect microvilli from physical or toxic injury or serve as a filter (Lehane, 1997). The villi may be somewhat shorter in the first int ring than in subsequent cells along the body axis (Sulston and Horvitz, 1977).
The intestinal cells are each very large and contain large nuclei with a prominent nucleolus, many mitochondria, extensive rough endoplasmic reticulum (RER), many ribosomes, and an extensive collection of membrane-bound vesicles and vacuoles. The nature of these organelles changes gradually as the animal ages. The digestive and metabolic activities of the intestine are central to the growth and development of the animal, and correspondingly, these organelles include yolk granules, recycling endosomes, autophagic vacuoles, and autofluorescent (gut) granules. Using light microscopy, some of these gut granules become visible as birefringent objects in older adults and are inferred to be secondary lysosomes involved in catabolism (Clokey and Jacobson, 1986).
The primary function of intestinal cells seems to be digestive because they secrete digestive enzymes (e.g. cysteine protease endodeoxyribonuclease) into the lumen and take up processed material and nutrients. The intestine also seems to be a large storage organ because it contains a large number of assorted storage granules that change in size, shape and number during development (White, 1988). In hermaphrodites, the intestine is also involved in synthesis and secretion of yolk material that is then transported to the oocytes through the body cavity (Kimble and Sharrock, 1983). The intestinal contents may also play role in miscellaneous functions carried out by nonintestinal cells in higher animals. For instance, the glycosyltransferases, which function in carbohydrate metabolism, comprise more than 70 genes in the C. elegans genome, and at least some appear to be expressed in the digestive tract (Griffitts et al., 2003; McKay et al., 2004). In addition, along with muscle, intestine is thought to be the major organ in which fatty acid metabolism takes place. Through the function of a glyoxylate cyclase (SRH-1) yolk fatty acid-derived acetylcoenzymeA is converted to succinate, from which carbohydrates are synthesized (Liu et al., 1995).
Anatomical and gene expression data both suggest that these functions differ along the length of the organ. For instance, the collection of membrane-bound organelles and vacuoles is more diverse and much more extensive in int rings I and II than further posterior (Borgonie et al., 1995). Without histochemical staining, it is still difficult to assign functions to each type of endosome, but open vacuoles of the anterior organ were proposed to release digestive enzymes into the gut lumen. In support of this observation, cysteine protease (CPR-1) expression is restricted to the anterior portions (among int rings I-VI) of the intestine (Britton et al., 1998). Yolk and lipid vacuoles predominate in posterior portions of the intestine, and these cells may be more active in nutrient and energy storage.
The posterior intestine also functions as the pacemaker of the defecation cycle. In C. elegans, defecation occurs in a rhythmic manner in tightly regulated cycles that are approximately 50 seconds long and have three distinct muscle contraction steps (see Alimentary system - Rectum). Inositol triphosphate (IP3) receptor-driven calcium oscillations in the posterior intestinal cells initiate the muscle contractions of the defecation cycle and the IP3 receptor is a central component of the timekeeping mechanism that regulates this behavioral rhythm (Dal Santo, 1999; see also Alimentary system - Rectum).
Recent studies show intestine functions as a cold temperature sensor (intestinal cells exhibit a robust increase in calcium level in response to cooling).This calcium response is greatly reduced in trpa-1 mutant worms, consistent with an important role for TRPA-1 in cold-reception in intestine and cold-dependent lifespan extension
(Xiao et al., 2013).
The intestine may change in shape and function rather dramatically in the dauer larva, which do not feed (Popham and Webster, 1979). The lumen becomes shrunken and the size and number of microvilli are greatly reduced (Albert and Riddle, 1988). When the animal emerges from the dauer state, these changes are reversed in the new L4 larva.
5 List of Intestinal Cells
1. First intestinal ring
int1DL
int1DR
int1VL
int1VR
2. Second intestinal ring
int2D
int2V
3. Third intestinal ring
int3D
int3D.a - postembryonic nuclear division
int3D.p - postembryonic nuclear division
int3V
int3V.a - postembryonic nuclear division
int3V.p - postembryonic nuclear division
4. Fourth intestinal ring
int4D
int4D.a - postembryonic nuclear division
int4D.p - postembryonic nuclear division
int4V
int4V.a - postembryonic nuclear division
int4V.p - postembryonic nuclear division
5. Fifth intestinal ring
int5L
int5L.a - postembryonic nuclear division
int5L.p - postembryonic nuclear division
int5R
int5R.a - postembryonic nuclear division
int5R.p - postembryonic nuclear division
6. Sixth intestinal ring
int6L
int6L.a - postembryonic nuclear division
int6L.p - postembryonic nuclear division
int6R
int6R.a - postembryonic nuclear division
int6R.p - postembryonic nuclear division
7. Seventh intestinal ring
int7L
int7L.a - postembryonic nuclear division
int7L.p - postembryonic nuclear division
int7R
int7R.a - postembryonic nuclear division
int7R.p - postembryonic nuclear division
8. Eighth intestinal ring
int8L
int8L.a - postembryonic nuclear division
int8L.p - postembryonic nuclear division
int8R
int8R.a - postembryonic nuclear division
int8R.p - postembryonic nuclear division
9. Ninth intestinal ring
int9L
int9L.a - postembryonic nuclear division
int9L.p - postembryonic nuclear division
int9R
int9R.a - postembryonic nuclear division
int9R.p - postembryonic nuclear division
6 References
Albert, P.S. and Riddle, D.L. 1988. Mutants of Caenorhabditis elegans that form dauer-like larvae. Dev. Biol. 126: 270-293. Abstract
Borgonie, G., Claeys, M., De Waele, D. and Coomans, A. 1995. Ultrastructure of the intestine of the bacteriophagous nematodes Caenorhabditis elegans, Panagrolaimus superbus and Acrobeloides maximus (Nematoda: Rhabditida). Fund. Appl. Nematol. 18: 123-133. Article
Britton, C., McKerrow, J.H. and Johnstone, I.L.1998. Regulation of the Caenorhabditis elegans gut cysteine protease gene cpr-1: Requirement for GATA motifs. J. Mol. Biol. 283: 15-27. Abstract
Bucher, E.A. and Seydoux, G.C. 1994. Gastrulation in the nematode Caenorhabditis elegans. Semin. Dev. Biol. 5: 121-130. Abstract
Clokey, G.V. and Jacobson, L.A. 1986. The autofluorescent "lipofuscin granules" in the intestinal cells of Caenorhabditis elegans are secondary lysosomes. Mech. Ageing Dev. 35: 79–94. Abstract
Dal Santo, P., Logan, M.A., Chisholm, A.D. and Jorgensen, E.M. 1999. The inositol triphosphate receptor regulates a 50-second behavioral rhythm in C. elegans. Cell 98: 757-767. Article
Goldstein, B. 1992. Induction of gut in Caenorhabditis elegans embryos. Nature 357: 255–257. Abstract
Goldstein, B. 1993. Establishment of gut fate in the E lineage of C. elegans: the roles of lineage-dependent mechanisms and cell interactions. Development 118: 1267–1277. Article
Graham, P.L., Johnson, J.J., Wang, S., Sibley, M.H., Gupta, M.C. and Kramer, J.M. 1997. Type IV collagen is detectable in most, but not all, basement membranes of Caenorhabditis elegans and assembles on tissues that do not express it. J. Cell. Biol. 137: 1171–1183. Article
Griffitts, J.S., Huffman, D.L., Whitacre, J.L., Barrows, B.D., Marroquin, L.D., Muller, R., Brown, J.R., Hennet, T., Esko, J.D. and Aroian, R.V. 2003. Resistance to a bacterial toxin is mediated by removal of a conserved glycosylation pathway required for toxin-host interactions. J. Biol. Chem. 278: 45594–45602. Article
Hedgecock, E.M. and White, J.G. 1985. Polyploid tissues in the nematode Caenorhabditis elegans. Dev. Biol. 107: 128–133. Abstract
Hermann, G.J., Leung, B., and Priess, J.R. 2000. Left-right asymmetry in C. elegans intestine organogenesis involves a LIN-12/Notch signaling pathway. Development 127: 3429–3440. Article
Huang, C.C., Hall, D.H., Hedgecock, E.M., Kao, G., Karantza, V., Vogel, B.E., Hutter, H., Chisholm, A.D., Yurchenco, P.D., and Wadsworth, W.G. 2003. Laminin alpha subunits and their role in C. elegans development. Development 130: 3343–3358. Article
Kang, S.H. and Kramer, J.M. 2000. Nidogen is nonessential and not required for normal type IV collagen localization in Caenorhabditis elegans. Mol. Biol. Cell 11: 3911–3923. Article
Kao, G., Huang, C.-C., Hedgecock, E.M., Hall, D.H. and Wadsworth, W. 2006. The role of the laminin β heterotrimer assembly and basement membrane function and development in C. elegans. Dev. Biol. 290: 211-219. Article
Kimble, J. and Sharrock, W. J. 1983. Tissue-specific synthesis of yolk proteins in Caenorhabditis elegans. Dev. Biol. 9: 189-196. Abstract
Labouesse, M. 2006. Epithelial junctions and attachments. WormBook, ed. The C. elegans Research Community, doi/10.1895/wormbook.1.56.1. Article
Lehane, M.J. 1997. Peritrophic matrix structure and function. Annu. Rev. Entomol. 42: 525–550. Abstract
Leung, B., Hermann, G.J. and Priess, J.R. 1999. Organogenesis of the Caenorhabditis elegans intestine. Dev. Biol. 216: 114–134. Article
Lin, R., Hill, R.J. and Priess, J.R. 1998. POP-1 and anterior-posterior fate decisions in C. elegans embryos. Cell 92: 229–239. Article
Lin, R., Thompson, S. and Priess, J.R. 1995. pop-1 encodes an Hmg box protein required for the specification of a mesoderm precursor in early C. elegans embryos. Cell 83: 599–609. Article
Liu, F., Thatcher, J.D., Barral, J.M. and Epstein, H.F. 1995. Bifunctional glyoxylate cycle protein of Caenorhabditis elegans: A developmentally regulated protein of intestine and muscle. Dev. Biol. 169: 399-414. Article
Maduro, M.F. and Rothman, J.H. 2002. Making worm guts: the gene regulatory network of the Caenorhabditis elegans endoderm. Dev. Biol. 246: 68–85. Article
McGhee J.D. 2007. The C. elegans intestine. In WormBook (ed. The C. elegans Research Community), WormBook, doi/10.1895/wormbook.1.133.1. Article
McKay, S.J., Johnsen, R., Khattra, J., Asano, J., Baillie, D.L., Chan, S., Dube, N., Fang, L., Goszczynski, B., Ha, E., Halfnight, E., Hollebakken, R., Huang, P., Hung, K., Jensen, V., Jones, S.J.M., Kai, H., Li, D., Mah, A., Marra, M., McGhee, J., Newbury, R., Pouzyrev, R., Riddle, D.L., Sonnhammer, E., Tian, H., Tu, D., Tyson, J.R., Vatcher, G., Warner, A., Wong, K., Zhao, Z. and Moerman, D.G. 2004. Gene expression profiling of cells, tissues and developmental stages of the nematode C. elegans. Cold Spring Harbor Symp. Quantit. Biol. 68: 159-69. Abstract
Mendenhall, A.R., Patricia M. Tedesco, P.M., Sands, B., Johnson, T.E. and Brent, R. 2015. Singlecell quantification of reporter geneexpression in live adult Caenorhabditis elegans reveals reproducible cell-specific expression patterns and underlying biological variation. PLoS ONE. 10: e0124289. Article
Nance, J., Lee, J.-Y. Goldstein, B. 2005. Gastrulation in C. elegans. In WormBook (ed. The C. elegans Research Community), WormBook, doi/10.1895/wormbook.1.23.1. Article
Pauli, F., Liu, Y., Kim, Y.A., Chen, P.-J. and Kim, S.K. 2006. Chromosomal clustering and GATA transcriptional regulation of intestine-expressed genes in C. elegans. Development 133: 287-295. Article
Popham, J.D. and Webster, J.M. 1979. Aspects of the fine structure of the dauer larva of the nematode C. elegans. Can. J. Zool. 57: 794-800. Abstract
Rocheleau, C.E., Yasuda, J., Shin, T-H., Lin, R. Sawa, H., Okano, H., Priess, J.R., Davis, R.J. and Mello, C.C. 1999. WRM-1 activates the LIT-1 protein kinase to transduce anterior posterior polarity signals in C. elegans. Cell 97: 717-726. Article
Schnabel, R. and Priess, J.R. 1997. Specification of Cell Fates in the Early Embryo. In C. elegans II (ed D.L. Riddle et al.). pp 361-382. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York. Article
Schulenburg, H., Kurz, C.L. and Ewbank, J.J. 2004. Evolution of the innate immune system: the worm. Immunol. Rev. 198: 36-58. Abstract
Sulston, J.E. and Horvitz, H.R. 1977. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev. Biol. 56: 110–156. Article
Sulston, J.E., Schierenberg, E., White, J.G. and Thomson, J.N. 1983. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100: 64-119. Article
Vogel, B.E. and Hedgecock, E.M. 2001. Hemicentin, a conserved extracellular member of the immunoglobulin superfamily, organizes epithelial and other cell attachments into oriented line-shaped junctions. Development 128: 883–894. Article
White J. 1988. The Anatomy. In The nematode C. elegans (ed. W.B. Wood) pp81-122. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York. Abstract
Wood, W.B., Bergmann, D. and Florance, A. 1996. Maternal effect of low temperature on handedness determination in C. elegans embryos. Dev. Genet. 19:222–230. Abstract
Xiao, R., Zhang, Bi., Dong, Y., Gong, J., Xu, T., Liu, J., Xu, X.Z.S. 2013. A genetic program promotesC. elegans longevity at cold temperatures via a thermosensitive TRP channel. Cell 152:806-817. Article
|