1 Overview
The alimentary system, one of the most complex portions of the nematode anatomy, is composed of a large variety of tissues and cell types (White, 1988). Direct intercellular connections between the alimentary tissues and the rest of the body are minimal. Topologically, this system forms a separate epithelial tube running inside the cylindrical body wall, separated from it by the pseudocoelomic space, and placed parallel to the gonad. At the anteriormost end, the pharyngeal epithelium connects to the arcade cells of the lips (See Interfacial Epithelial Cells). The posteriormost region of the alimentary tract, the rectal epithelium and anus, is surrounded by the tail hypodermis and the dorsorectal ganglion. One pair of somatic neurons (RIPL/RIPR) penetrates the pharyngeal basal lamina to make gap junctions with a pair of pharyngeal neurons (I1L/R). Otherwise the only connection that spans the two systems (somatic vs. enteric) are the four specialized enteric muscle cells that act on the intestine and rectal valve while being driven by the somatic motor neurons, AVL and DVB.
The alimentary system is divided into the foregut (stomodeum; buccal cavity and the pharynx), the midgut (intestine), and the hindgut (proctodeum; rectum and anus in hermaphrodites and cloaca in males) and contains a total of 127 cells (AlimFIG 1). The stomodeum and proctodeum are lined with cuticle, which is shed along with body cuticle during molts (See Cuticle) (Bird and Bird, 1991). The stomodeum and proctodeum are also regions into which enteric glands open; pharyngeal glands open to the stomodeum and rectal glands open to the proctodeum. Ingested material flows through the digestive tract by the muscular pumping and peristalsis of the pharynx at the anterior end, and the waste material is discarded through the opening of the anus at the posterior end by the action of the enteric muscles. The intestine itself is devoid of any muscular structure in C. elegans, although its rear portion can be contracted by the stomatointestinal (SI) muscles (see Intestine). During defecation, the body wall muscles also contribute to the control of internal pressure and concentration of the gut contents before the expulsion of the waste material (see Rectum and Anus). Developmentally, the intestine is endodermal in origin, whereas the stomodeum and proctodeum have a mixed lineage from ectodermal and mesodermal origins (Bird and Bird, 1991).
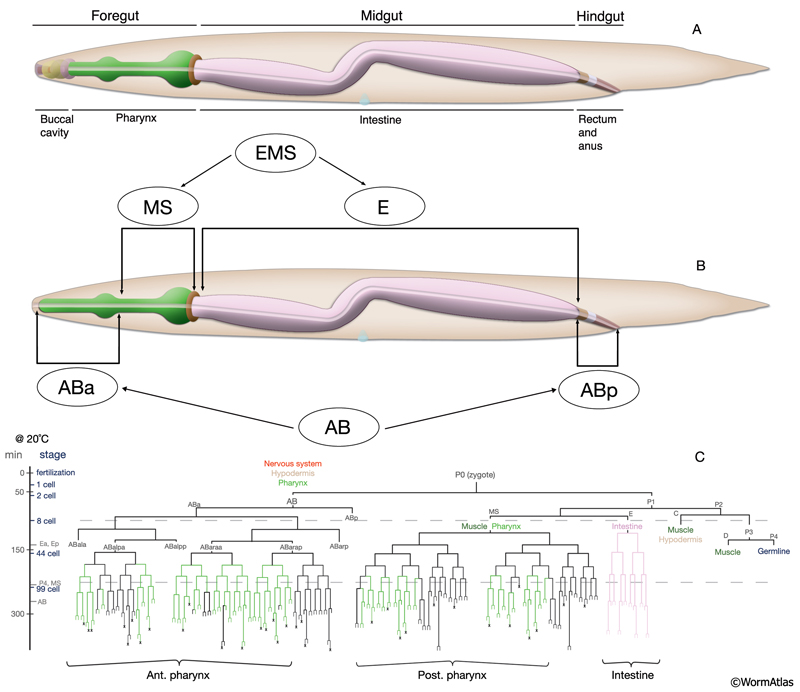 The alimentary system is described in different sections by its individual organs along the length of the body, from anterior to posterior (see Pharynx, Intestine and Rectum and Anus sections). The tissues that connect to the pharyngeal epithelium and are more anterior to the pharyngeal epithelium within the lips (hyp1, hyp2, hyp3, the lip sensilla, and the arcade cells) are described in the Epithelial System - Hypodermis. |
2 References
Bird, A. F. and Bird, J. 1991. The structure of nematodes. Academic Press, California.
White, J. 1988. The Anatomy. In The nematode C. elegans (ed. W. B. Wood). Chapter 4. pp 81-122. Cold Spring Harbor Laboratory Press, New York. Abstract
Fukushige, T., Schroeder, D.F., Allen, F.L., Goszczynski, B. and McGhee, J.D. 1996. Modulation of gene expression in the embryonic digestive tract of C. elegans. Dev. Biol. 178: 276-288. Abstract
|
